Here's an example of a matrix (corresponding to the graph with black edges only), and its Cholesky factorization. We triangulate the graph by any pairs of neighbors of v which have higher index than v, for every v. Extra edges added in this way correspond to extra non-zero entries in Cholesky decomposition of the original matrix. Different color edges/matrix entries correspond to non-zero entries in Cholesky factorization not present in original matrix
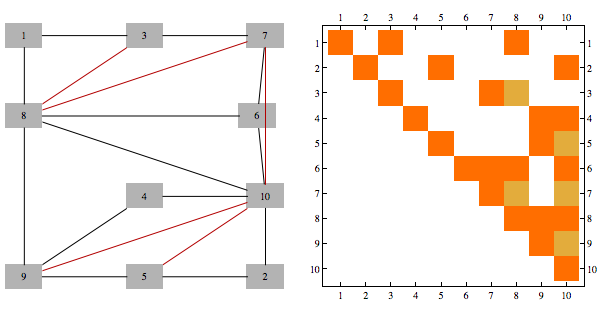
Furthermore, junction tree we get using min-fill heuristic of a graph is same structure as what is used to perform Cholesky factorization using parallel multi-frontal method
Here's a chordal matrix in perfect order and its Cholesky factorization
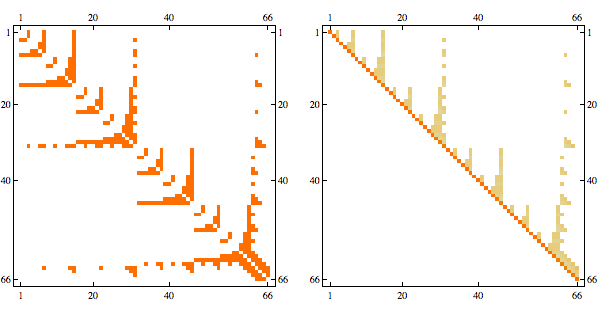
The fact that Cholesky factorization has the same sparsity structure as the original matrix confirms that its a chordal matrix in perfect elimination order
Here's the graph representation of the matrix
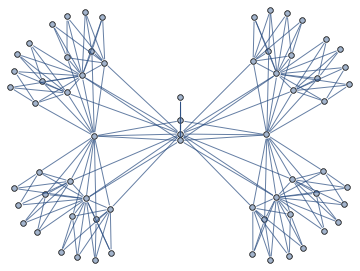
Junction tree decomposition of this graph reveals the following clique structure
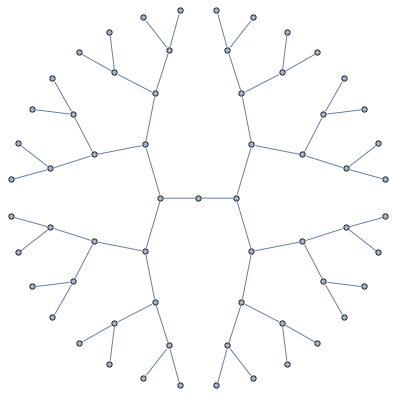
High branching structure of decomposition means that Cholesky factorization can be highly parallelized, in particular, the very last step of Cholesky decomposition of this matrix in minfill order consists of 32 independent tasks that can be done in parallel.
I packaged together some Mathematica code for these kinds of tasks
20 comments:
THANKS FOR THE INFORMATION...
<a href="http://www.chloros.in/digital-marketing-internship.htmlhttp://www.chloros.in/digital-marketing-internship.html>Digital Marketing Internship Program in Bangalore</a>
Excellent machine learning blog,thanks for sharing...
Seo Internship in Bangalore
Smo Internship in Bangalore
Digital Marketing Internship Program in Bangalore
PERMAINAN ONLINE TERBESAR DI INDONESIA
Website paling ternama dan paling terpercaya di Asia ^^
Sistem pelayanan 24 Jam Non-Stop bersama dengan CS Berpengalaman respon tercepat :)
Memiliki 8 Jenis game yang sangat digemari oleh seluruh peminat poker / domino
- Adu Q
- Bandar Q
- Bandar Sakong
- Bandar Poker
- Poker
- Domino 99
- Capsa Susun
- BANDAR66 / ADU BALAK ( GAME TERBARU )
Permainan Judi online yang menggunakan uang asli dan mendapatkan uang asli ^^
* Minimal Deposit : 20.000
* Minimal Withdraw : 20.000
* Deposit dan Withdraw 24 jam Non stop ( Kecuali Bank offline / gangguan )
* Bonus REFFERAL 15 % Seumur hidup tanpa syarat
* Bonus ROLLINGAN 0.3 % Dibagikan 5 hari 1 kali
* Proses Deposit & Withdraw PALING CEPAT
* Sistem keamanan Terbaru & Terjamin
* Poker Online Terpercaya
* Live chat yang Responsive
* Support lebih banyak bank LOKAL
Contact Us
Website : sahabat2019.org
WA 1 : +85515769793
WA 2 : +855972076840
Telegram 1 :+85515769793
Telegram 2 : +855972076840
LINE : SAHABATQQ
Kami Siap Melayani anda 24 jam Nonstop
Permainan Sabung Ayam Online di Agen BOLAVITA , dengan minimal deposit hanya Rp 25.000 saja , dan minimal betting hanya Rp 10.000 saja sudah bisa mainkan permainan Sabung Ayam
http://agensabungayam.logdown.com/post/7858081-kasta-bulu-ayam-bangkok-yang-indah-dan-ditakuti
Produk Kami Judi Sabung Ayam Online S128, SV388 & KungfuChiken.
https://www.sateayam.vip/
https://m1.hj128.vip/
Daftar Sabung Ayam sv388
Daftar Sabung Ayam Online S128
Agen Sabung Ayam Online Bolavita Banyak Bonus dan Promo Mari Bergabung :
Promo Sabung Ayam Terbaru 8x Win Beruntun.
Bolavita Bisa Deposit Via OVO & GO-Pay.
Sabung Ayam Deposit Via Pulsa XL & TSEL 25rb.
Promo Promo BOLAVITA
Telegram : +62812-2222-995
Wechat : Bolavita
WA : +62812-2222-995
Line : cs_bolavita
PERMAINAN ONLINE TERBESAR DI INDONESIA
Website paling ternama dan paling terpercaya di Asia ^^
Sistem pelayanan 24 Jam Non-Stop bersama dengan CS Berpengalaman respon tercepat :)
Memiliki 8 Jenis game yang sangat digemari oleh seluruh peminat poker / domino
- Adu Q
- Bandar Q
- Bandar Sakong
- Bandar Poker
- Poker
- Domino 99
- Capsa Susun
- BANDAR66 / ADU BALAK ( GAME TERBARU )
Permainan Judi online yang menggunakan uang asli dan mendapatkan uang asli ^^
* Minimal Deposit : 20.000
* Minimal Withdraw : 20.000
* Deposit dan Withdraw 24 jam Non stop ( Kecuali Bank offline / gangguan )
* Bonus REFFERAL 15 % Seumur hidup tanpa syarat
* Bonus ROLLINGAN 0.3 % Dibagikan 5 hari 1 kali
* Proses Deposit & Withdraw PALING CEPAT
* Sistem keamanan Terbaru & Terjamin
* Poker Online Terpercaya
* Live chat yang Responsive
* Support lebih banyak bank LOKAL
Contact Us
Website : sahabat2019.org
WA 1 : +85515769793
WA 2 : +855972076840
Telegram 1 :+85515769793
Telegram 2 : +855972076840
LINE : SAHABATQQ
Kami Siap Melayani anda 24 jam Nonstop
Saya memiliki skor kredit yang sangat rendah sehingga upaya saya untuk meminjam dari Bank ditolak. Saya bangkrut sampai-sampai saya tidak mampu makan tiga kali sehari dan juga saya benar-benar bangkrut karena nama saya identik dengan kemiskinan. saya berhutang baik dari teman-teman saya dan juga dari rentenir hidup saya di bawah ancaman saya harus melarikan diri dari rumah dan saya membawa anak-anak saya untuk bertemu ibu mertua saya karena sifat ancaman yang saya terima dari orang-orang yang meminjamkan saya uang Jadi saya harus mencari cara cepat dan mendesak untuk membayar kembali uang itu dan juga memulai bisnis baru usaha pertama saya sangat mengerikan karena saya ditipu sebesar Rp5.390.020.00 saya harus pindah juga dua minggu kemudian saya kehilangan Rp350.000,00 kepada pemberi pinjaman yang curang jadi saya turun secara finansial dan emosional karena ini adalah yang paling tidak saya harapkan sehingga seorang teman saya memberi tahu saya untuk menghubungi email ini: :( iskandalestari.kreditpersatuan@gmail.com) bahwa saya harus meminta jumlah berapa pun berharap agar Bunda Iskandar selalu menjadi kembali untuk memberikan bantuan keuangan kepada siapa pun yang membutuhkan sehingga saya meminta untuk jumlah Rp850.000.000,00 dalam waktu 24 jam cerita saya berubah untuk selamanya saya membayar semua hutang saya dan saya juga memiliki cukup uang untuk membiayai sendiri bisnis semua terima kasih kepada teman saya yang memperkenalkan saya kepada ibu khususnya dan juga kepada Ibu Iskandar pada umumnya untuk mengubah rasa malu saya menjadi terkenal
Atas perkenan: ISKANDAR LESTARI LOAN COMPANY
Email: (iskandalestari.kreditpersatuan@gmail.com)
Nice article
Im very support the owner
Agen Domino99
SahabatQQ
Bandar poker Terpercaya
Bandar poker Terpercaya
Bandar poker Terpercaya
Bandar poker Terpercaya
Bandar poker Terpercaya
Bandar poker Terpercaya
Dewacintaqq
Dewacintaqq.com
Dewa cintaqq
Poker Online
Domino 99
BandarQ
link alternatif Dewacintaqq
Daftar Dewacintaqq
Bandar poker Terpercaya
dewacintaqq
dewacintaqq.com
dewa cintaqq
Cinta
daftar dewacintaqq
link alternatif dewacintaqq
Bandar poker Terpercaya
Bandar poker Terpercaya
SITUS TOGEL ONLINE
Situs Togel Hongkong
Situs Togel Singapore
Bandar poker terpercaya
Daftar Dominohalo
http://45.76.123.180/?ref=Nisa93
Bandar poker terpercaya
Daftar twinpoker88
http://149.28.170.64/?ref=twinpoker21
Bandar poker terpercaya
Daftar Delimapoker
http://45.76.112.124/?ref=nisa2122
Bandar poker terpercaya
Daftar Papadomino
DewacintaQQ
link alternatif Dewacintaqq
Deposit
Deposit dewacintaqq
Poker Indonesia
Poker Online Indonesia
Bandar QQ
Agen Ceme Online
Daftar Situs Poker
Poker Online Terpercaya
Judi Poker
Poker Online Uang Asli
Poker Uang Asli
Situs Poker Online
Agen Poker
Poker88
Capsa Susun Online
daftar idn poker
idnpoker
bigceme
ceme
ceme online
bandar ceme
idn poker
Idnplay
idn play
poker
poker idn
agen poker online
poker online
situs poker online
judi online
situs judi online
Papadomino
Delimapoker
Twinpoker88
Dominohalo
Dewacintaqq
dewacintaqq.
Bandar poker terpercaya
situs judi online
judi online
situs poker online
poker online
dominoqq
domino99
bandarq
-----------------------------------------------------------------------------
Bandarq online terpercaya
Delimapoker situs resmi daftar judi online
bandar66
sakong
capsa
domino99
aduq uang asli qq online domino QQ
bandarq,judi online
bandar poker
qq online
---------------------------------------------------------------------------------
Bandar poker terpercaya
judi online
situs judi online
poker online
dominoqq
bandarqq
bandar poker
bandarq
sakong
aduq
capsa susun
------------------------------------------------------------------------------------
Bandar poker terpercaya
judi poker online
bandar poker online
judi online
poker online
dominoqq
situs judi online
situs poker online
------------------------------------------------------------------------------------
Bandar poker terpercaya
Poker Indonesia
Poker Online Indonesia
Agen Ceme Online
Daftar Situs Poker
Poker Online Terpercaya
Judi Poker
Poker Online Uang Asli
Poker Uang Asli
Situs Poker Online
Agen Poker
Situs Judi Online Terpercaya
bandar poker
situs judi
poker online
ceme online
bandar ceme
situs poker
bandar judi
poker idn
judi poker
judi online
agen judi
bandar judi
poker idn
agen poker
idn poker
Situs Judi Poker Online
DominoQQ IDN POKER
Agen judi kartu deposit uang asli pkv games
poker online
dominoqq
bandarqq
bandar poker
bandarq
sakong
aduq
capsa susun
Perang bacarat
Bandar poker Terpercaya
Bandar poker Terpercaya
Bandar poker Terpercaya
Bandar poker Terpercaya
Bandar poker Terpercaya
Bandar poker Terpercaya
Dewacintaqq
Dewacintaqq.com
Dewa cintaqq
Poker Online
Domino 99
BandarQ
link alternatif Dewacintaqq
Daftar Dewacintaqq
Bandar poker Terpercaya
dewacintaqq
dewacintaqq.com
dewa cintaqq
Cinta
daftar dewacintaqq
link alternatif dewacintaqq
Bandar poker Terpercaya
Bandar poker Terpercaya
SITUS TOGEL ONLINE
Situs Togel Hongkong
Situs Togel Singapore
Bandar poker terpercaya
Daftar Dominohalo
Bandar poker terpercaya
Daftar twinpoker88
Bandar poker terpercaya
Daftar Delimapoker
Bandar poker terpercaya
Daftar Papadomino
DewacintaQQ
link alternatif Dewacintaqq
Deposit
Deposit dewacintaqq
Poker Indonesia
Poker Online Indonesia
Bandar QQ
Agen Ceme Online
Daftar Situs Poker
Poker Online Terpercaya
Judi Poker
Poker Online Uang Asli
Poker Uang Asli
Situs Poker Online
Agen Poker
Poker88
Capsa Susun Online
daftar idn poker
idnpoker
bigceme
ceme
ceme online
bandar ceme
idn poker
Idnplay
idn play
poker
poker idn
agen poker online
poker online
situs poker online
judi online
situs judi online
papadomino
papadomino.com
papa domino
daftar papadomino
link alternatif papadomino
http://45.32.190.103/?ref=vivi293
delimapoker
delimapoker.com
delima poker
daftar delimapoker
link alternatif delimapoker
http://45.76.112.124/?ref=nisa2122
twinpoker88
twin poker88
twinpoker88.com
daftar delimapoker
Link alternatif twinpoker88
http://149.28.170.64/?ref=twinpoker21
dominohalo
dominohalo.com
domino halo
daftar dominohalo
link alternatif dominohalo
http://45.76.123.180/?ref=Nisa93
bandar poker terpercaya
bandar togel terpercaya
papadomino
papadomino.com
papa domino
daftar papadomino
link alternatif papadomino
http://45.32.190.103/?ref=vivi293
papadomino
papadomino.com
papa domino
daftar papadomino
link alternatif papadomino
delimapoker
delimapoker.com
delima poker
daftar delimapoker
link alternatif delimapoker
http://45.76.112.124/?ref=nisa2122
delimapoker
delimapoker.com
delima poker
daftar delimapoker
link alternatif delimapoker
twinpoker88
twin poker88
twinpoker88.com
daftar delimapoker
Link alternatif twinpoker88
http://149.28.170.64/?ref=twinpoker21
twinpoker88
twin poker88
twinpoker88.com
daftar delimapoker
Link alternatif twinpoker88
dominohalo
dominohalo.com
domino halo
daftar dominohalo
link alternatif dominohalo
http://45.76.123.180/?ref=Nisa93
dominohalo
dominohalo.com
domino halo
daftar dominohalo
link alternatif dominohalo
http://45.76.123.180/?ref=Nisa93
dewacintaqq
dewacintaqq.com
dewa cintaqq
daftar dewacintaqq
link alternatif dewacintaqq
Poker Indonesia
Poker Online Indonesia
Bandar QQ
Agen Ceme Online
Daftar Situs Poker
Poker Online Terpercaya
Judi Poker
Poker Online Uang Asli
Poker Uang Asli
Situs Poker Online
Agen Poker
poker online
Capsa Susun Online
situs poker online
situs judi online
Poker Indonesia
Poker Online Indonesia
Bandar QQ
Agen Ceme Online
Daftar Situs Poker
Poker Online Terpercaya
Judi Poker
Poker Online Uang Asli
Poker Uang Asli
Situs Poker Online
Agen Poker
poker online
Capsa Susun Online
situs poker online
situs judi online
Saya memiliki skor kredit yang sangat rendah sehingga upaya saya untuk meminjam dari Bank ditolak. Saya bangkrut sampai-sampai saya tidak mampu makan tiga kali sehari dan juga saya benar-benar bangkrut karena nama saya identik dengan kemiskinan. saya berhutang baik dari teman-teman saya dan juga dari rentenir hidup saya di bawah ancaman saya harus melarikan diri dari rumah dan saya membawa anak-anak saya untuk bertemu ibu mertua saya karena sifat ancaman yang saya terima dari orang-orang yang meminjamkan saya uang Jadi saya harus mencari cara cepat dan mendesak untuk membayar kembali uang itu dan juga memulai bisnis baru usaha pertama saya sangat mengerikan karena saya ditipu sebesar Rp5.390.020.00 saya harus pindah juga dua minggu kemudian saya kehilangan Rp350.000,00 kepada pemberi pinjaman yang curang jadi saya turun secara finansial dan emosional karena ini adalah yang paling tidak saya harapkan sehingga seorang teman saya memberi tahu saya untuk menghubungi email ini: :( iskandalestari.kreditpersatuan@gmail.com) bahwa saya harus meminta jumlah berapa pun berharap agar Bunda Iskandar selalu menjadi kembali untuk memberikan bantuan keuangan kepada siapa pun yang membutuhkan sehingga saya meminta untuk jumlah Rp850.000.000,00 dalam waktu 24 jam cerita saya berubah untuk selamanya saya membayar semua hutang saya dan saya juga memiliki cukup uang untuk membiayai sendiri bisnis semua terima kasih kepada teman saya yang memperkenalkan saya kepada ibu khususnya dan juga kepada Ibu Iskandar pada umumnya untuk mengubah rasa malu saya menjadi terkenal
Atas perkenan: ISKANDAR LESTARI LOAN COMPANY
Email: (iskandalestari.kreditpersatuan@gmail.com)
Situs Online Terbesar, Terlengkap Dan Terpercaya
Aslijudi
Prediksi Parlay
NowGoal
Link Alternati Aslijudi
Kunjungi Juga Blog Kami Lainnya
Aslislotonline.blogspot.com
Aslijudibola.blogspot.com
Aslilivecasino.blogspot.com
Aslisituspokeronline.blogspot.com
Aslisabungayam.blogspot.com
gotrainings - Right Choice For A Bright Future. For more details visit below link.
machine learning course in hyderabad
best machine learning course in india
Thanks for being truthful, I will immediately share your blog on my Facebook and Instagram platform to many other members of my group and also take the advantage to share with you the new
UNDETECTED FAKE BIRTH CERTIFICATE FOREIGNER USE TO OBTAIN PASSPORTS ABROAD FREELY which many are ignorant of at the moment, it is also the easiest way to obtain dual nationality through this birth certificate in any country. I request your permission to join our 379.3k members TELEGRAM GROUP
to share with us your daily post for it to also reach my audience. Thanks, I am Scott from Globex, we await you on our platform too.
You can call me Scott.
meta777
daftar meta777
link meta777
daftar meta777
meta777
daftar meta777
daftar meta777
link alternatif meta777
My response to your website is very good, so I can learn a lot on judi online, with so many websites like togel online I can learn from your website I am very thankful
agen togeland join naw situs togel
I was diagnosed with Parkinson’s disease four years ago. After traditional medications stopped working, I tried a herbal treatment from NaturePath Herbal Clinic Within months, my tremors eased, balance improved, and I regained my energy. It’s been life-changing I feel like myself again. If you or a loved one has Parkinson’s, I recommend checking out their natural approach at [www.naturepathherbalclinic.com]. info@naturepathherbalclinic.com
Post a Comment