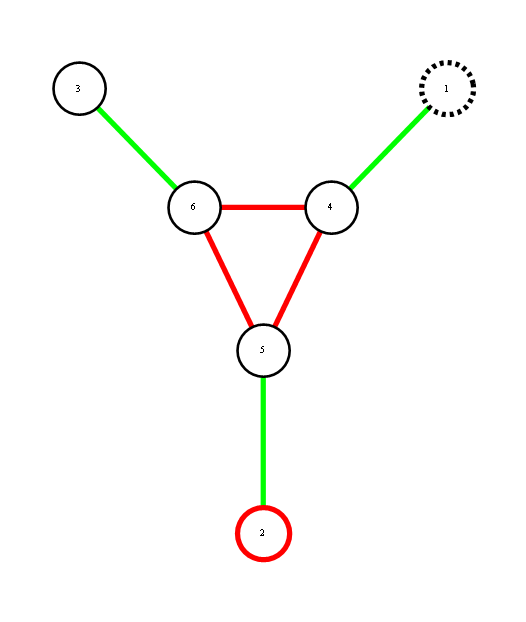
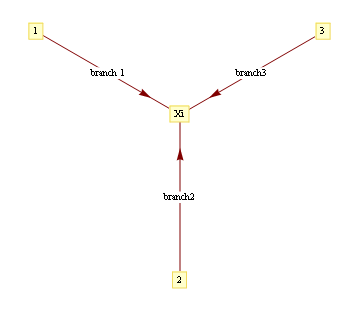
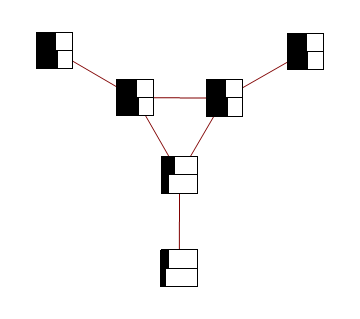
Red represents -1 potential, green represents +1 potential. More precisely, it's the diagram of the following model, where x's are {-1,1}-valued
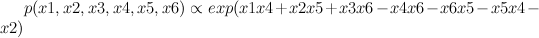
We'd like to find the odds of x1 being 1. You could do it by dividing sum of potentials over labellings with x1=1 by corresponding sum over labellings with x1=-1.
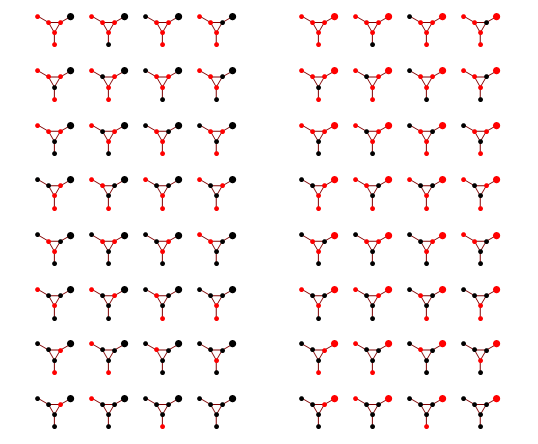
In a graph without cycles, such sums can be computed in time linear in the number of nodes by means of dynamic programming. In such cases it reduces to a series of local updates
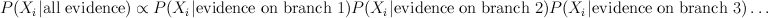
where evidence (or potential) from each branch is found by marginalizing over the separating variable
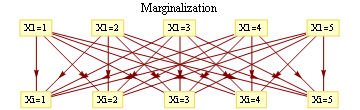
For binary states, update equations become particularly simple.
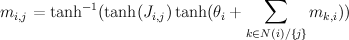
Here, m_{i,j} is the "message" being sent from node i to node j. If we view message passing as a way of computing marginal conditioned on the evidence, then message {i,j} represents the log-odds that node i is positive, given all the evidence on the other side of of the edge i,j. If our graph has loops, then "other side" of the edge doesn't make sense, but you can use the equations anyway, and this is called "loopy BP".
For binary networks with at most 1 loop, Yair Weiss showed that even though loopy BP may give incorrect probability estimates, highest probability labels under BP will be the same.
Implementation of loopy BP is below. Basically you add up all the messages from neighbours, subtract the message of the target node, then "squash" the message towards zero using ArcTanh[Tanh[J]Tanh[x]] function. Procedure loopyBP takes a matrix of connections, local potential strengths, and returns a function that represents one iteration of BP. Below you see the first few iterations for the graphical model above


To get the final potential at any given node, just add up all the incoming messages. For the graph above, we can plot the probabilities and see that they fall on the correct side of zero after 10 iterations, so that is much smaller computational burden than exact marginalization.
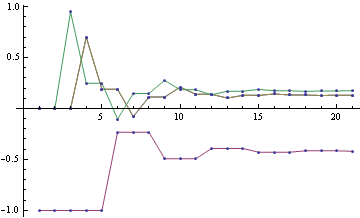
Below you can see the estimates produced by running loopy BP for 200 iterations. The top bar at each node is the actual probability, the bottom bar is the estimated probability using loopy BP. You can see that the feedback makes the estimates overly confident.
For graphs with more than one cycle, loopy BP is no longer guaranteed to converge, or to give MAP labelling when it converges. For instance, here's an animation of loopy BP applied to square Ising lattice with randomly initialized neighbour potentials
Some methods like the Convex-Concave procedure and Generalized Belief Propagation improve on BP's convergence and accuracy.
Mathematica notebook used to produce these figures
11 comments:
Great one. I am not sure what the commands graph2mat and symmetrize do. Do we need some additional libraries? or is it that, the script would not run as it is with mathematica 7. It would be nice if you could please comment on this. This motivates me to make a BP decoding demo. Will update you on that. It seems that the Mathematica 7 does not recognize graph2mat for instance!
In all, I enjoyed reading your blog. I also liked the one formula for prime numbers! Great show.
Ooops, forgot about graph2mat, it's here
http://yaroslavvb.com/research/ideas/loopy-blogpost/utils.nb
Just execute the initialization cells in that notebook and you should be set. Let me know if you run into any other problems running the notebook
Thanks Yaroslav for the update. It seems to be fine now. Meanwhile I found a workaround using AdjacencyMatrix command. Now anyway your initialization cell make it easier.
Best
R
I like this topic.
sexy girls in London
Excellent machine learning blog,thanks for sharing...
Seo Internship in Bangalore
Smo Internship in Bangalore
Digital Marketing Internship Program in Bangalore
바카라사이트
파워볼사이트
카지노사이트
파워볼사이트
This information is worth everyone's attention.
Interesting problem! Understanding variable dependencies in probabilistic models can be tricky. Visualizing the potentials with colors definitely helps. It reminds me of strategically placing toppings in Papa's Pizzeria to maximize customer happiness, but with probabilities instead of pepperoni! Solving for the odds of x1 being 1 requires careful calculation of the sums of potentials across all possible configurations, a bit like optimizing your pizza-making process.
´´´´´████████´´´´
´´`´███▒▒▒▒███´´´´´
´´´███▒●▒▒●▒██´´´
´´´███▒▒▒▒▒▒██´´´´´
´´´███▒▒💋▒▒██´
´´██████▒▒███´´´´´
´██████▒▒▒▒███´´
██████▒▒▒▒▒▒███´´´´
´´▓▓▓▓▓▓▓▓▓▓▓▓▓▒´´
´´▒▒▒▒▓▓▓▓▓▓▓▓▓▒´´´´´
´.▒▒▒´´▓▓▓▓▓▓▓▓▒´´´´´
´.▒▒´´´´▓▓▓▓▓▓▓▒
..▒▒.´´´´▓▓▓▓▓▓▓▒
´▒▒▒▒▒▒▒▒▒▒▒▒
´´´´´´´´´███████´´´´
´´´´´´´´████████´´´´´´
´´´´´´´█████████´´´´´
´´´´´´██████████´´´
´´´´´´██████████´´
´´´´´´´█████████´
´´´´´´´█████████´
´´´´´´´´████████´´´
________▒▒▒▒▒
_________▒▒▒▒
_________▒▒▒▒
________▒▒_▒▒
_______▒▒__▒▒
_____ ▒▒___▒▒
_____▒▒___▒▒
____▒▒____▒▒
___▒▒_____▒▒
_███______▒▒
█_███____███
█__███_ _█_███
█ ███
Wow this is awesome. i love it.
I really enjoyed reading this post, I always appreciate topics like this being discussed to us. Information very nice. I will follow post Thanks for sharing.
So nice article, i found it very explanatory and informative, keep doing this nice job!
This is a great post! Really keep up the good work!
Post a Comment